 Organizers:
Organizers:
Maike Weber, Sven Berres, Svitlana Rozanova, Dirk Winkelhardt, Karin Schork (Ruhr University Bochum, BioInfra.Prot, de.NBI)
Trainers/Speakers (in no specific order):
Timo Sachsenberg, Samuel Wein (CIBI / University of Tübingen, Germany), Robert Heyer (BiGi / Bielefeld University & ISAS Dortmund, Germany,), Daniel Wibberg (CAU / Forschungszentrum Jülich GmbH, Germany), Magnus Palmblad (Leiden University Medical Center, Netherlands), Louise Buur (University of Applied Sciences Upper Austria, Austria), Veit Schwämmle (University of Southern Denmark, Denmark), Marie Locard-Paulet (Université de Toulouse, France), Pietro Crivello (University Hospital Essen, Germany), Mathias Wilhelm (Technical University of Munich, Germany), Wout Bittremieux (University of Antwerp, Belgium), Ralf Gabriels (Ghent University, Belgium), Laurent Gatto (UCLouvain, Belgium)
Date:
07.09.2026 (Starting at 12:00 CEST) – 11.09.2026 (Ending at 15:00 CEST)
Location:
Bochum, Germany – Ruhr Universität Bochum, Beckmannshof
Contents:
The de.NBI Spring School 2026 focuses on bioinformatics for the analysis of mass-spectrometry-based peptide and protein data.
Participants will cover data formats, peptide identification, protein inference,and quantification, gaining an understanding of how each compontent of the bottom-up proteomics works and how everything fits together. In addition the limits and pitfalls of proteomics data and analysis will be covered to ensure each participant is able to take precautions in their future work.
Advanced topics like machine learning in proteomics, immunopeptidomics, single-cell proteomics, and metaproteomics, will also be introduced.
The program combines lectures and hands-on sessions and is aimed at PhD students, postdoctoral researchers, and scientists with a background in bioinformatics and molecular biology.
Participants will gain the skills needed to analyze and interpret proteomics data in their own research.
We are happy to share that there are no registration fees for the event. To fully support your stay, we will provide accommodation and local travel tickets within Bochum. Furthermore, meals (breakfast / lunch) and social events are included in the program, so you can focus entirely on the workshop and networking.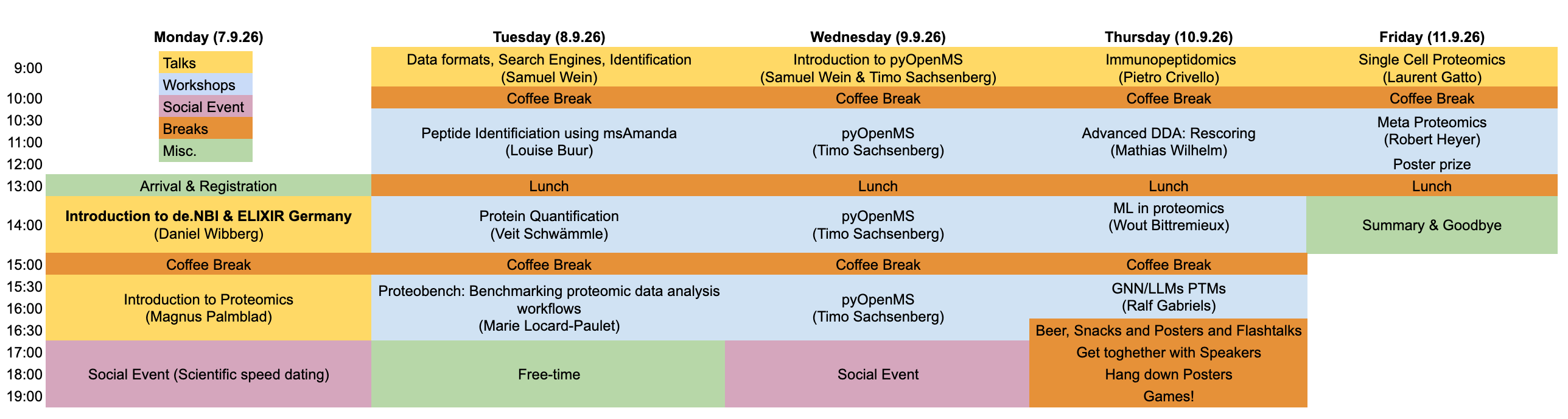
Prerequisites:
The Summer School is targeted towards early career scientists like PhD students, but also experienced scientists, who want to learn more about Bioinformatics for Proteomics. A basic understanding of Python programming is required.
Keywords:
Proteomics, Mass Spectrometry, Data Analysis, DDA & DIA
Tools:
msAmanda, Proteobench, pyOpenMS
Contact:
Application:
https://events.hifis.net/event/3935/
Deadline: 01.06.2026

